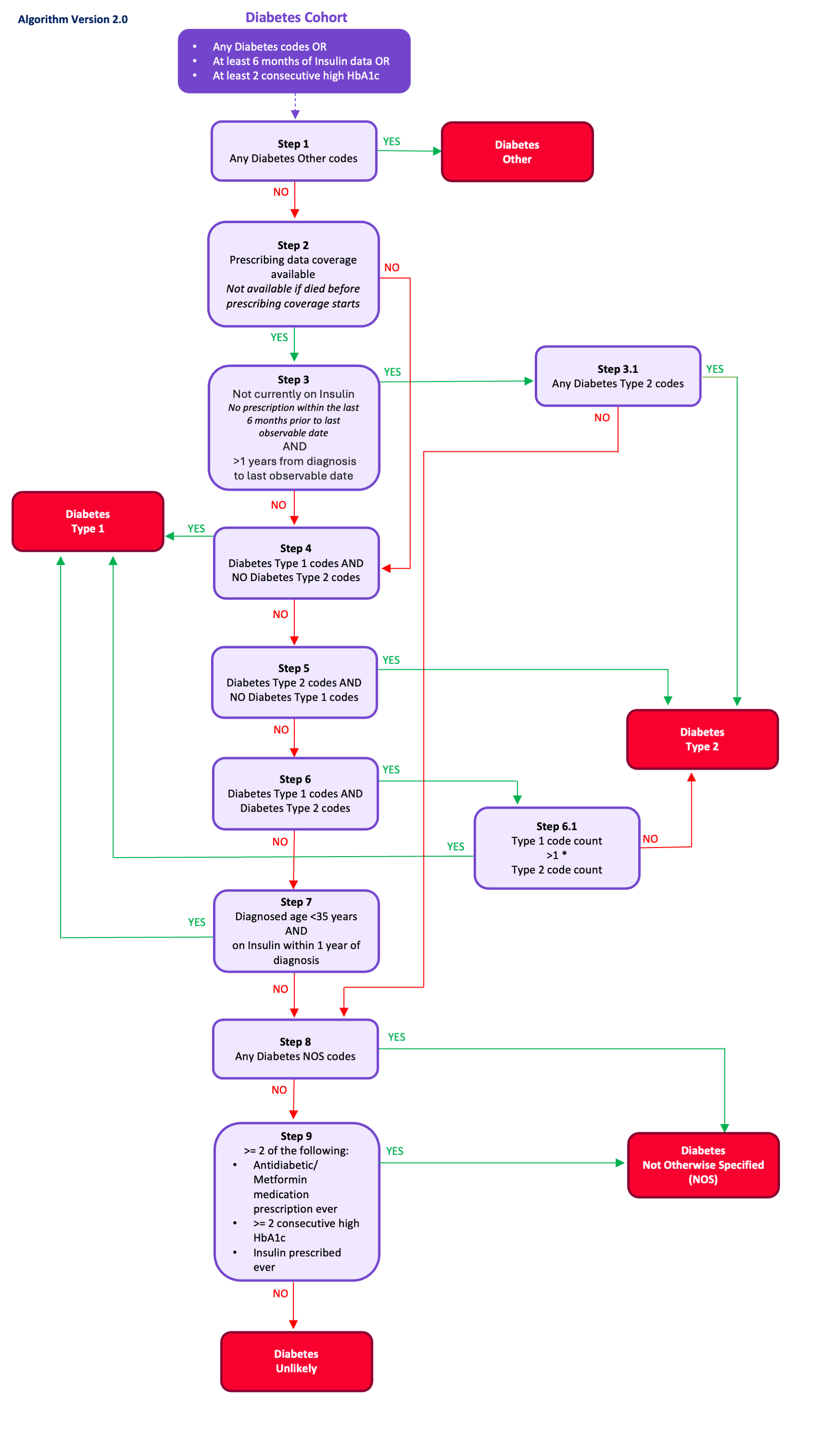
Diabetes
PH4062 / 9273
Chalmers, F, Young K, Walker E, Chilala C, Coombe A, Shields B, Preiss D, Cezard G, MacArthur J, Dennis J, Valabhji J, Kaptoge S, Khunti K, Mafham M, Rutter MK, Lessels S, Shi W, Wild S, Eastwood S, Denaxas S, Bolton T, Thomas N, Pearson E
Apr 23, 2026
Overview
Project NameDiabetes Data Science CatalystPhenotype TypeDisease or syndromeSexBothValid Event Date RangeNo dataCoding SystemSNOMED CT codesBNF codesICD10 codesData SourcesOntologyNo dataCollectionsBHF Data Science CentreTagsNo dataDefinition
Definition
Clinical Trials
No Trials Endorsements
No endorsement Implementation
Implementation
Clinical Codelist
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
PUBLISHED
Publication
Related publications
No known publicationsCitation Example
API
To Export Phenotype Details:
Format API JSON site_root/api/v1/phenotypes/PH4062/version/9273/detail/?format=json R Package library(ConceptLibraryClient)
# Connect to API
client = ConceptLibraryClient::Connection$new(public=TRUE)
# Get details of Phenotype
phenotype_details = client$phenotypes$get_detail(
'PH4062',
version_id=9273
)Py Package from pyconceptlibraryclient import Client
# Connect to API
client = Client(public=True)
# Get details of Phenotype
phenotype_detail = client.phenotypes.get_detail(
'PH4062',
version_id=9273
)To Export Phenotype Code List:
Format API JSON site_root/api/v1/phenotypes/PH4062/version/9273/export/codes/?format=json R Package library(ConceptLibraryClient)
# Connect to API
client = ConceptLibraryClient::Connection$new(public=TRUE)
# Get codelist of Phenotype
phenotype_codelist = client$phenotypes$get_codelist(
'PH4062',
version_id=9273
)Py Package from pyconceptlibraryclient import Client
# Connect to API
client = Client(public=True)
# Get codelist of Phenotype
phenotype_codelist = client.phenotypes.get_codelist(
'PH4062',
version_id=9273
)Version History