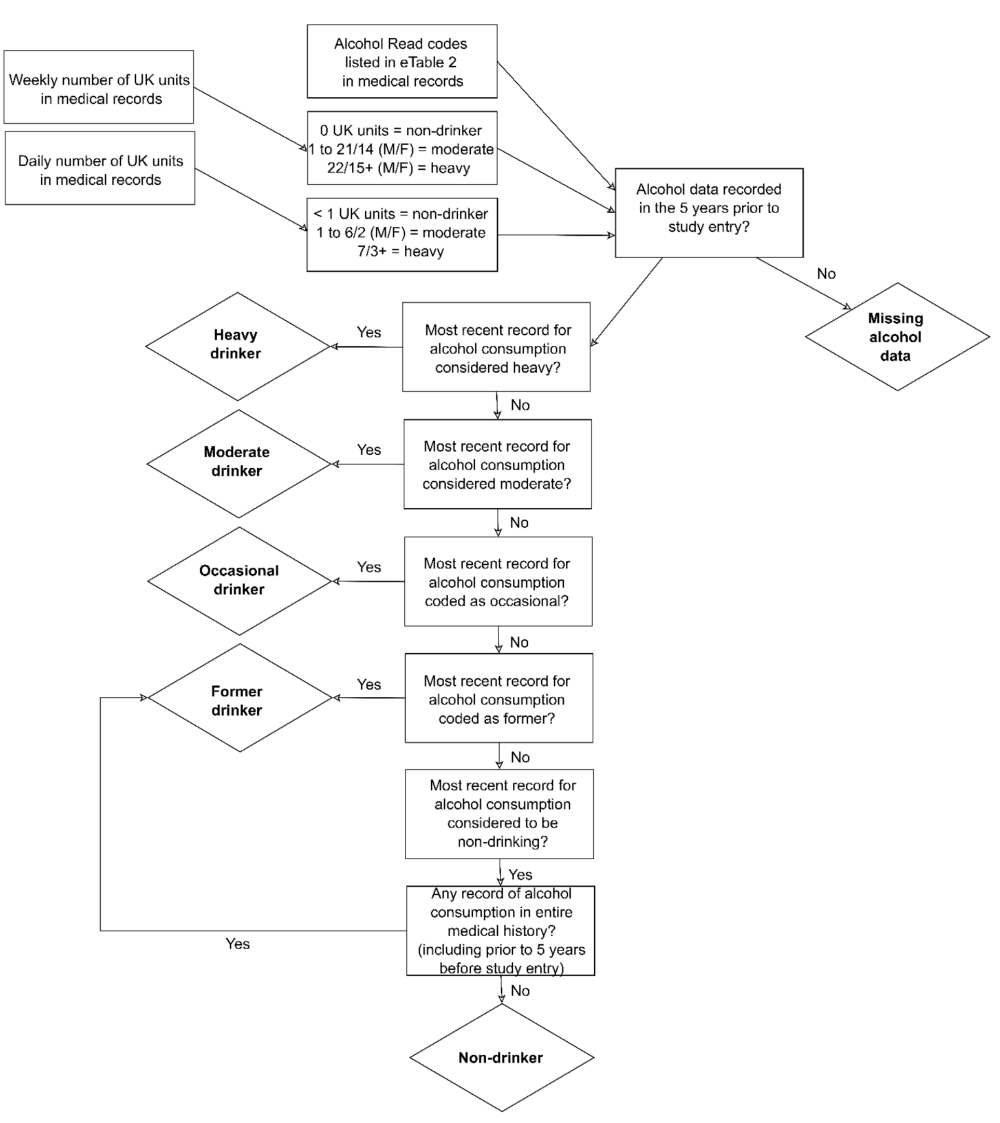
Alcohol Consumption
PH9 / 1510
Bell S, Daskalopoulou M, Rapsomaniki E, George J, Britton A, Bobak M, Casas JP, Dale CE, Denaxas S, Shah AD, Hemingway H
Oct 19, 2021
Overview
Phenotype TypeLifestyle risk factorSexBothValid Event Date Range01/01/1999 - 01/07/2016Coding SystemRead codes v2Data SourcesCollectionsPhenotype LibraryTagsNo dataOntologyNo dataDefinition
Implementation
Phenoflow ID
Please see the PhenoFLOW website here for more information on workflow-based Phenotypes.
No dataImplementation
Clinical Codelist
PUBLISHED
PUBLISHED
PUBLISHED
Publication
Related publications
Gho JMIH et al. An electronic health records cohort study on heart failure following myocardial infarction in England: incidence and predictors. BMJ Open. 2018 Mar 3;8(3):e018331. doi: 10.1136/bmjopen-2017-018331. PMID: 29502083
(DOI: 10.1136/bmjopen-2017-018331)Steele AJ et al. Machine learning models in electronic health records can outperform conventional survival models for predicting patient mortality in coronary artery disease. PLoS One. 2018 Aug 31;13(8):e0202344. doi: 10.1371/journal.pone.0202344. eCollection 2018. PMID: 30169498
(DOI: 10.1371/journal.pone.0202344)Archangelidi O et al. Clinically recorded heart rate and incidence of 12 coronary, cardiac, cerebrovascular and peripheral arterial diseases in 233,970 men and women: A linked electronic health record study. Eur J Prev Cardiol. 2018 Sep;25(14):1485-1495. doi: 10.1177/2047487318785228. Epub 2018 Jul 2. PMID: 29966429
(DOI: 10.1177/2047487318785228)Koudstaal S et al. Prognostic burden of heart failure recorded in primary care, acute hospital admissions, or both: a population-based linked electronic health record cohort study in 2.1 million people. Eur J Heart Fail. 2017 Sep;19(9):1119-1127. doi: 10.1002/ejhf.709. Epub 2016 Dec 23. PMID: 28008698
(DOI: 10.1002/ejhf.709)Chung SC et al. Time spent at blood pressure target and the risk of death and cardiovascular diseases. PLoS One. 2018 Sep 5;13(9):e0202359. doi: 10.1371/journal.pone.0202359. eCollection 2018. PMID: 30183734
(DOI: 10.1371/journal.pone.0202359)Bell S et al. Association between clinically recorded alcohol consumption and initial presentation of 12 cardiovascular diseases: population based cohort study using linked health records. BMJ. 2017 Mar 22;356:j909. PMID: 28331015
Pasea L et al. Personalising the decision for prolonged dual antiplatelet therapy: development, validation and potential impact of prognostic models for cardiovascular events and bleeding in myocardial infarction survivors. Eur Heart J. 2017 Apr 7;38(14):1048-1055. doi: 10.1093/eurheartj/ehw683. PMID: 28329300
(DOI: 10.1093/eurheartj/ehw683)Shah AD et al. Neutrophil Counts and Initial Presentation of 12 Cardiovascular Diseases: A CALIBER Cohort Study. J Am Coll Cardiol. 2017 Mar 7;69(9):1160-1169. doi: 10.1016/j.jacc.2016.12.022. PMID: 28254179
(DOI: 10.1016/j.jacc.2016.12.022)Asaria M et al. Using electronic health records to predict costs and outcomes in stable coronary artery disease. Heart. 2016 May 15;102(10):755-62. doi: 10.1136/heartjnl-2015-308850. Epub 2016 Feb 10. PMID: 26864674
(DOI: 10.1136/heartjnl-2015-308850)Daskalopoulou M et al. Depression as a Risk Factor for the Initial Presentation of Twelve Cardiac, Cerebrovascular, and Peripheral Arterial Diseases: Data Linkage Study of 1.9 Million Women and Men. PLoS One. 2016 Apr 22;11(4):e0153838. doi: 10.1371/journal.pone.0153838. eCollection 2016. PMID: 27105076
(DOI: 10.1371/journal.pone.0153838)Pujades-Rodriguez M et al. Associations between polymyalgia rheumatica and giant cell arteritis and 12 cardiovascular diseases. Heart. 2016 Mar;102(5):383-9. doi: 10.1136/heartjnl-2015-308514. Epub 2016 Jan 19. PMID: 26786818
(DOI: 10.1136/heartjnl-2015-308514)Pujades-Rodriguez M et al. Rheumatoid Arthritis and Incidence of Twelve Initial Presentations of Cardiovascular Disease: A Population Record-Linkage Cohort Study in England. PLoS One. 2016 Mar 15;11(3):e0151245. doi: 10.1371/journal.pone.0151245. eCollection 2016. PMID: 26978266
(DOI: 10.1371/journal.pone.0151245)Shah AD et al. Low eosinophil and low lymphocyte counts and the incidence of 12 cardiovascular diseases: a CALIBER cohort study. Open Heart. 2016 Sep 5;3(2):e000477. doi: 10.1136/openhrt-2016-000477. eCollection 2016. PMID: 27621833
(DOI: 10.1136/openhrt-2016-000477)Timmis A et al. Prolonged dual antiplatelet therapy in stable coronary disease: comparative observational study of benefits and harms in unselected versus trial populations. BMJ. 2016 Jun 22;353:i3163. PMID: 27334486
Walker S et al. Long-term healthcare use and costs in patients with stable coronary artery disease: a population-based cohort using linked health records (CALIBER). Eur Heart J Qual Care Clin Outcomes. 2016 Jan 20;2(2):125-140. doi: 10.1093/ehjqcco/qcw003. PMID: 27042338
(DOI: 10.1093/ehjqcco/qcw003)George J et al. How Does Cardiovascular Disease First Present in Women and Men? Incidence of 12 Cardiovascular Diseases in a Contemporary Cohort of 1,937,360 People. Circulation. 2015 Oct 6;132(14):1320-8. doi: 10.1161/CIRCULATIONAHA.114.013797. Epub 2015 Sep 1. PMID: 26330414
(DOI: 10.1161/CIRCULATIONAHA.114.013797)Morley KI et al. Defining disease phenotypes using national linked electronic health records: a case study of atrial fibrillation. PLoS One. 2014 Nov 4;9(11):e110900. doi: 10.1371/journal.pone.0110900. eCollection 2014. PMID: 25369203
(DOI: 10.1371/journal.pone.0110900)Pujades-Rodriguez M et al. Heterogeneous associations between smoking and a wide range of initial presentations of cardiovascular disease in 1937360 people in England: lifetime risks and implications for risk prediction. Int J Epidemiol. 2015 Feb;44(1):129-41. doi: 10.1093/ije/dyu218. Epub 2014 Nov 20. PMID: 25416721
(DOI: 10.1093/ije/dyu218)Pujades-Rodriguez M et al. Socioeconomic deprivation and the incidence of 12 cardiovascular diseases in 1.9 million women and men: implications for risk prediction and prevention. PLoS One. 2014 Aug 21;9(8):e104671. doi: 10.1371/journal.pone.0104671. eCollection 2014. PMID: 25144739
(DOI: 10.1371/journal.pone.0104671)Rapsomaniki E et al. Blood pressure and incidence of twelve cardiovascular diseases: lifetime risks, healthy life-years lost, and age-specific associations in 1.25 million people. Lancet. 2014 May 31;383(9932):1899-911. doi: 10.1016/S0140-6736(14)60685-1. PMID: 24881994
(DOI: 10.1016/S0140-6736(14)60685-1)Shah AD et al. Type 2 diabetes and incidence of cardiovascular diseases: a cohort study in 1.9 million people. Lancet Diabetes Endocrinol. 2015 Feb;3(2):105-13. doi: 10.1016/S2213-8587(14)70219-0. Epub 2014 Nov 11. PMID: 25466521
(DOI: 10.1016/S2213-8587(14)70219-0)Rapsomaniki E et al. Prognostic models for stable coronary artery disease based on electronic health record cohort of 102 023 patients. Eur Heart J. 2014 Apr;35(13):844-52. doi: 10.1093/eurheartj/eht533. Epub 2013 Dec 17. PMID: 24353280
(DOI: 10.1093/eurheartj/eht533)
Citation Example
API
To Export Phenotype Details:
Format API JSON site_root/api/v1/phenotypes/PH9/version/1510/detail/?format=json R Package library(ConceptLibraryClient)
# Connect to API
client = ConceptLibraryClient::Connection$new(public=TRUE)
# Get details of Phenotype
phenotype_details = client$phenotypes$get_detail(
'PH9',
version_id=1510
)Py Package from pyconceptlibraryclient import Client
# Connect to API
client = Client(public=True)
# Get details of Phenotype
phenotype_detail = client.phenotypes.get_detail(
'PH9',
version_id=1510
)To Export Phenotype Code List:
Format API JSON site_root/api/v1/phenotypes/PH9/version/1510/export/codes/?format=json R Package library(ConceptLibraryClient)
# Connect to API
client = ConceptLibraryClient::Connection$new(public=TRUE)
# Get codelist of Phenotype
phenotype_codelist = client$phenotypes$get_codelist(
'PH9',
version_id=1510
)Py Package from pyconceptlibraryclient import Client
# Connect to API
client = Client(public=True)
# Get codelist of Phenotype
phenotype_codelist = client.phenotypes.get_codelist(
'PH9',
version_id=1510
)Version History